Charalampos (Babis) Tsourakakis@CMU
Dynamic Programming & aCGH Denoising
Dynamic Programming & aCGH Denoising
Approximation Algorithms for Speeding up
Dynamic Programming and Denoising aCGH data
About this Web Site
This web site contains the code, links to the datasets and a brief summary of our contributions contained in our submission to the Journal of Experimental Algorithmics.Contributions
Let P=(P1,..,Pn) the noisy input measurements. Our contributions are can be summarized in the following bullets:- We propose a new formulation of the aCGH denoising problem which we solve using a vanilla dynamic programming algorithm in O(n^2) time.
- We provide a technique which approximates the optimal value of our objective function within additive \epsilon error and runs in Õ(n^{4/3+\delta}log(\frac{U}{\epsilon}) time, where \delta is an arbitrarily small positive constant and U=max(sqrt{C},(|P_i|)_{i=1,...,n}).
- We provide another technique for approximate dynamic programming which solves the corresponding recurrence within a multiplicative factor of (1+$\epsilon$) and runs in $O(n \log{n} / \epsilon )$.
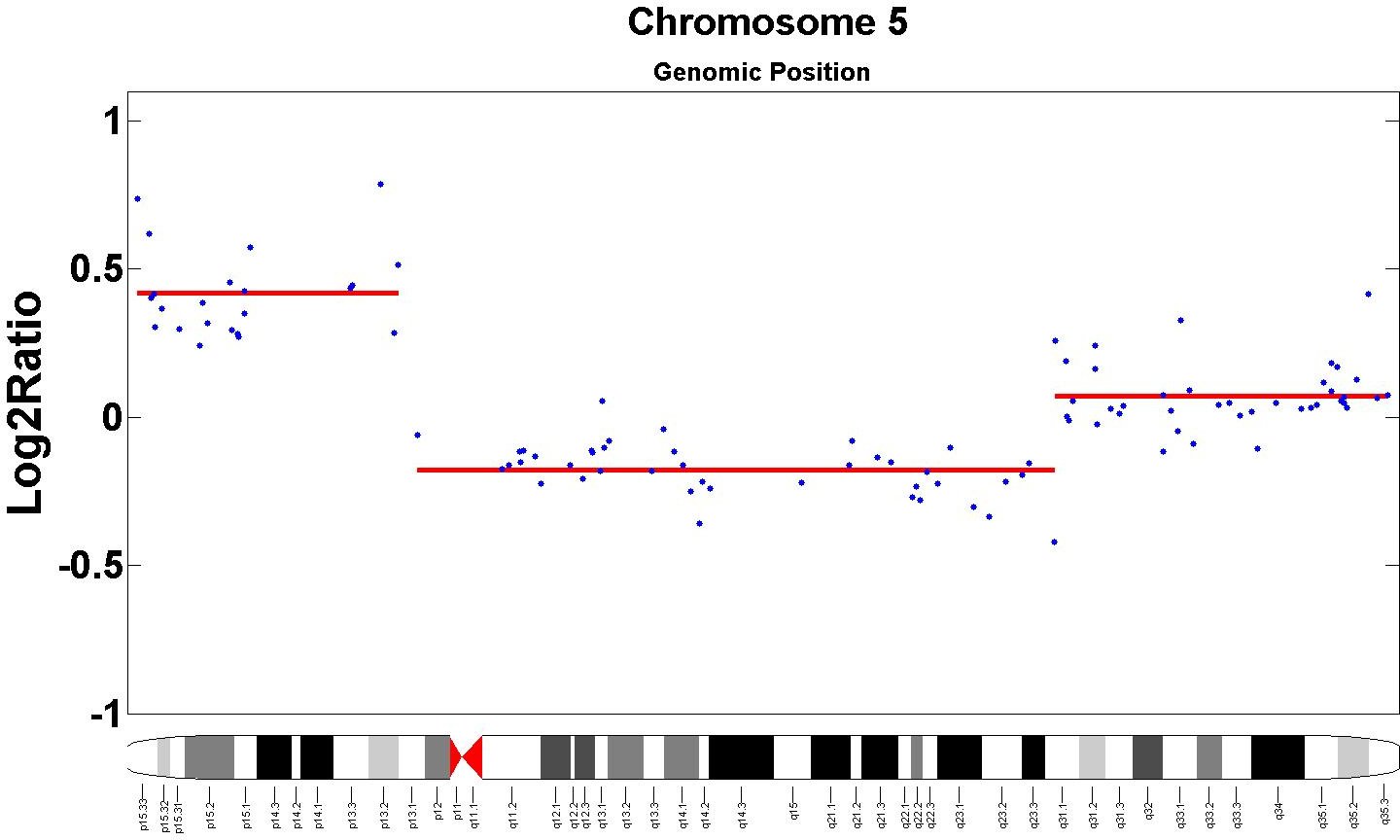
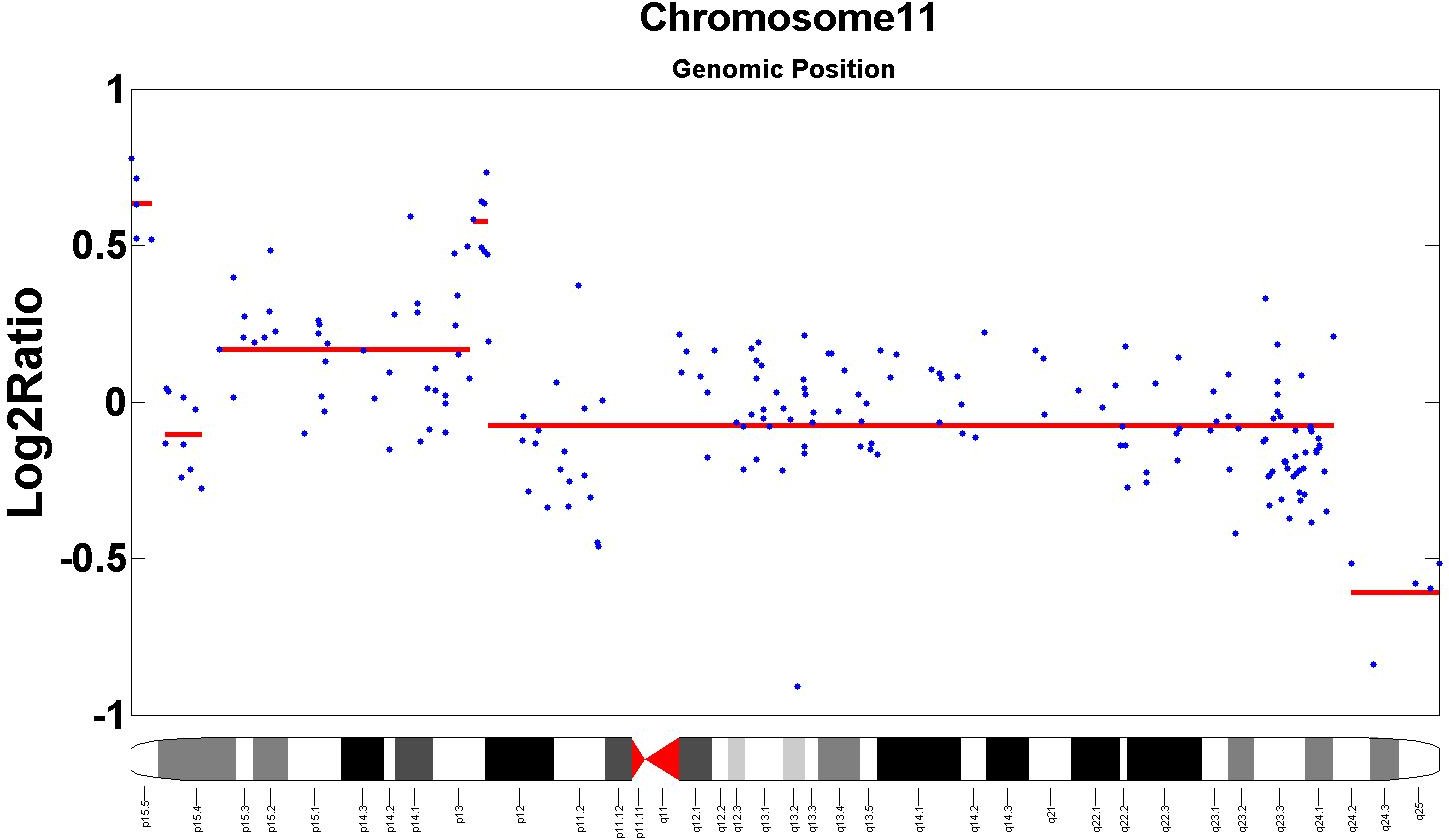
- We validate our proposed model on both synthetic and real data. Specifically, our segmentations result in superior precision and recall compared to leading competitors on benchmarks of synthetic data and real data from the Coriell cell lines. In addition, we are able to find several novel markers not recorded in the benchmarks but supported in the oncology literature.
Related Publications
- Approximation Algorithms for Speeding up Dynamic Programming and Denoising aCGH data
with Richard Peng, Maria Tsiarli,Russell Schwartz and Gary L. Miller
ACM Journal of Experimental Algorithmics (JEA 2011)
- Approximate Dynamic Programming using Halfspace Queries and Multiscale Monge decomposition
with Gary L. Miller, Richard Peng and Russell Schwartz
Symposium on Discrete Algorithms (SODA 2011)
Source Code
CGHTRIMMER is implemented in MATLAB. You can download the source code from this link (zip) or from this link (tar.gz). A C implementation (not used in the paper) with a simple demo is also available from this link (.c).Datasets
| Dataset | Availability |
|---|---|
| Lai et al 2005 | http://compbio.med.harvard.edu |
| Willenbrock et al. 2005 | http://www.cbs.dtu.dk/~hanni/aCGH/ |
| Coriell Cell Lines | load coriell_baccgh in Matlab or Nature |
| Berkeley Breast Cancer | See bundled dataset .mat inside our code or ICBP.LBL.GOV |